Folding (and then stapling) DNA Origami
The study of DNA is a topic most people are introduced to during their time at school and is often described using the phrase ‘the code of life’. This code is made up of the individual bases known as adenine, cytosine, guanine and thymine and, depending on their order, contain the information to create and build up cells into any living being. To do so, the bases combine into the iconic double helical structure of DNA which was definitively imaged via X-ray crystallography nearly 70 years ago in 1953.
By the early 2000s, scientists had started to move away from considering DNA as just genetic material and, presumably realising its robust structure made it suitable for manipulation, moved into creating a new area of science called DNA Origami. This area came into its own, in both technique and name, in 2006, with the release of a Nature paper detailing a new method of using DNA as a biomaterial to create novel and interesting shapes such as tiles, stars and even smiley faces. This really brought to the forefront the idea of DNA progressing beyond being only genetic material and, with this new technique, opened up an accessible and relatively straightforward method to create structures, which are all made of nothing but DNA.
A chemical process which people would be most familiar with would use the concept of a series of chemical reactions using reagents to produce a final product. DNA Origami also has ‘reagents’ but these are one long piece of single-stranded DNA, called a scaffold, and many smaller strands called staples. Once these have been prepared, the process is that of self-assembly using the technique of annealing (heating up and letting the solution cool down) where staples pinch together different areas of the long scaffold to create a DNA Origami - the single stranded staples and scaffold combine to produce a structure comprised of double helices.
Experimentally, the structures are visualised through imaging techniques such as atomic force microscopy (AFM) and transmission electron microscopy (TEM). However, computer simulations are often used to model the DNA to help provide more detail than may be possible through the aforementioned experimental methods. Such work can be carried out using the Oxford-developed software oxDNA which is a versatile molecular dynamics package and can be used to simulate the structure and analyse the properties of many systems including DNA Origami. Research on both the simulation and synthetic sides of this field is ongoing by members of the Doye, Louis and Turberfield groups in the chemistry and physics departments at the University of Oxford. Combining experiment and simulation provides a powerful approach to design and realise new Origamis with novel structures and functionalities; thus the field provides a good example of the value of a successful collaboration – an example being the work recently published on the topic of DNA T-junctions (Young et al., 2020).
Although some of the structures are used as fun demonstratives of the capabilities of the technique, the work undertaken has greatly helped to expand the knowledge of the action, behaviour and predictability of DNA and, with ever more sophisticated methods of easily designing DNA Origami being developed and released, this will surely lead to more groups exploring its capabilities and uses.
Through innovative areas like this, it is possible to look beyond DNA as just a genetic code and move into new areas which make use of it as a biomodelling material. It is truly a marvel how far the study of DNA has progressed in just 70 years and, with scientists now starting to build DNA machines, it is stunning to think about other uses it may have by the end of the next 70.
Article by Hannah Fowler
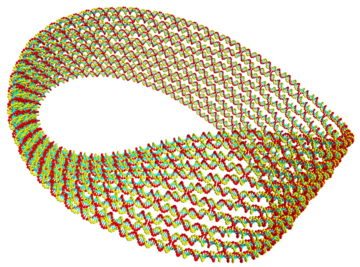

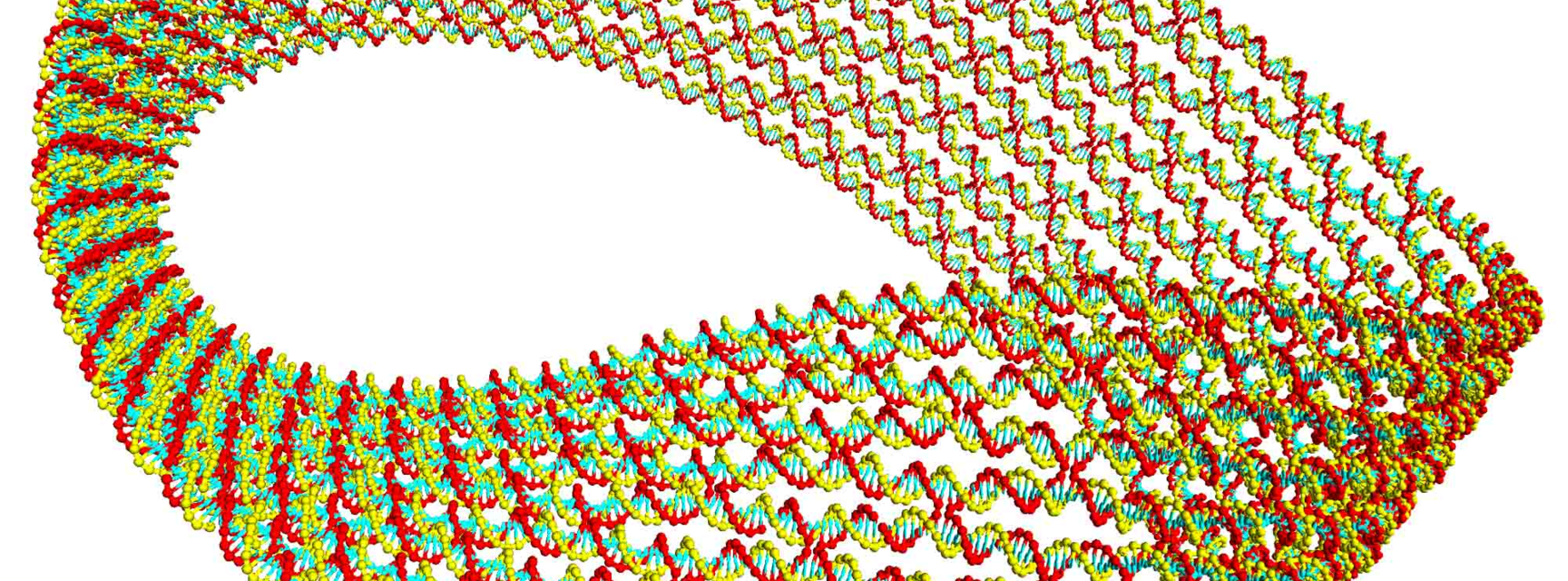
Image: The simulation result, from using oxDNA, of a DNA Origami Möbius strip: the scaffold strand iterates around the whole structure and shorter staple strands hold the scaffold in a folded state. Based on the experimental design from Han et al. Nature Nanotechnology, 2010, 5, 712.
References: P. W. Rothemund, Nature, 2006, 440, 297; J. P. K. Doye et al., Phys. Chem. Chem. Phys., 2013, 15, 20395; K. G. Young et al., Angew. Chem. Int., 2020, 59, 15942; Chemistry World article on DNA machines.